In a recent study published in Emerging Infectious Diseases, researchers described the evolution of the mpox virus (MPXV) before the 2022 outbreak.

Background
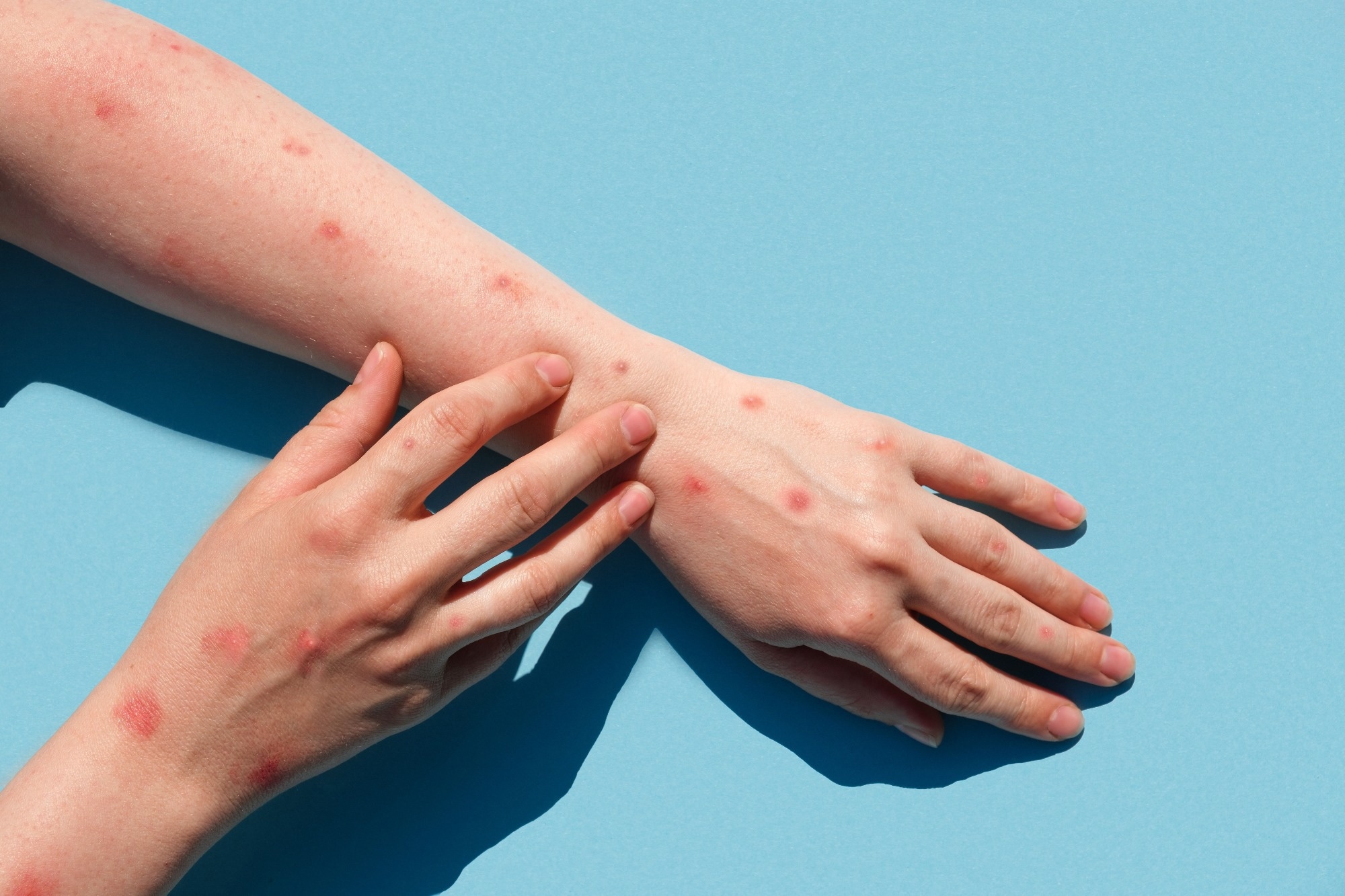
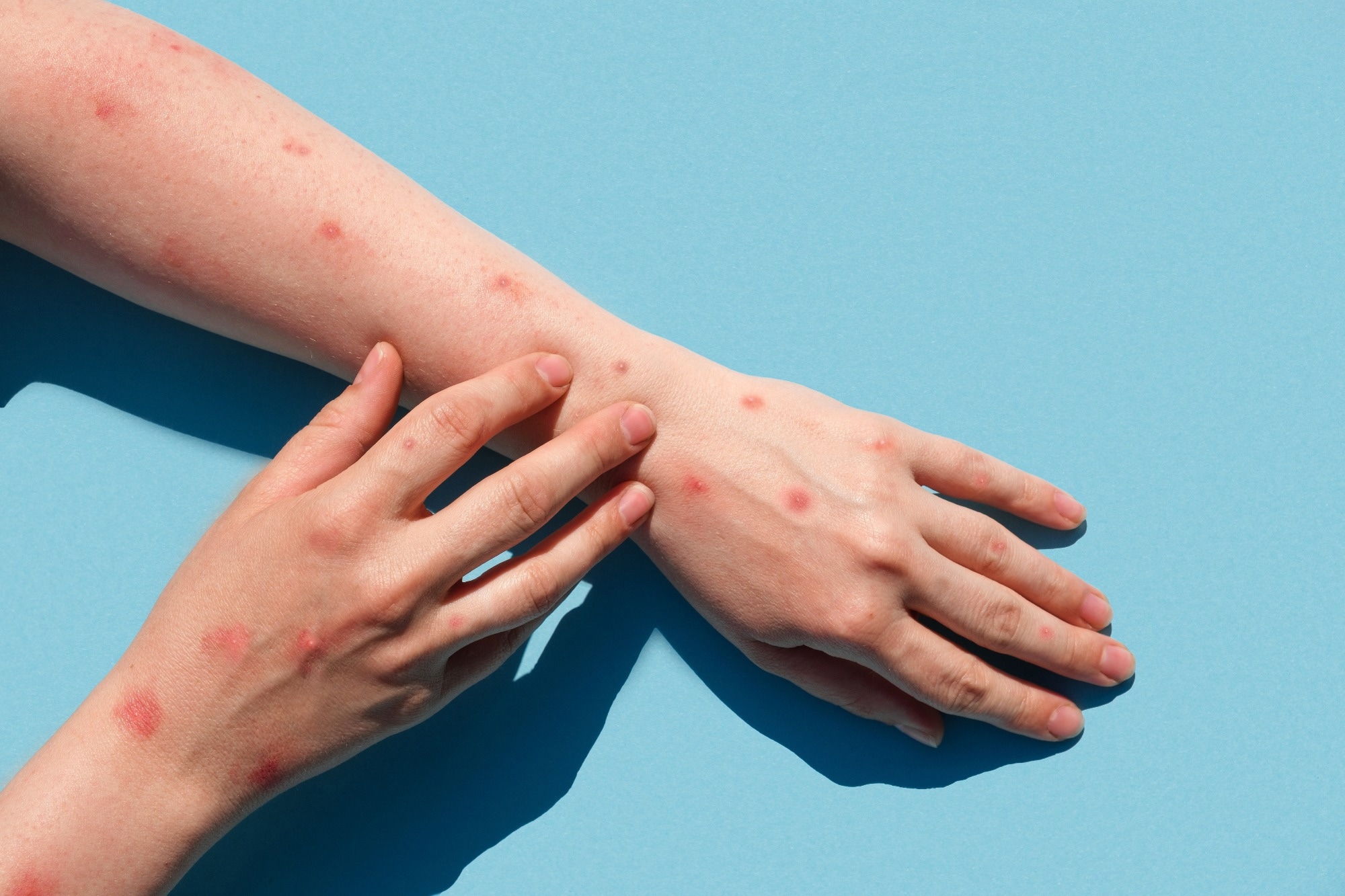
Mpox virus is a double-stranded deoxyribonucleic acid (DNA) virus that has resulted in outbreaks that are often brief and self-limiting due to inefficient human transmission. Initial epidemiologic investigations demonstrated continuous transmission from human to human in non-endemic European nations via intimate contacts, such as sexual networks.
Portugal announced the first MPXV genome sequence from the epidemic on 19 May 2022, and many more sequences that give information on viral transmission are available. Early phylogenetic analysis suggested that the virus responsible for the 2022 MPXV outbreak was from the MPXV clade II, which is found to have less severity than clade I. This suggested that the recent MPXV introduction into communities from non-MPXV-endemic nations led to the outbreak. Nonetheless, extensive research is needed to determine the exact evolutionary process of MPXV.
Phylogenetic investigation of MPXV
Phylogenetic analysis of almost 105 MPXV genomes indicated that 2022 viruses belonged to two clades that may be linked back to the last epidemic in 2017–2018. All 2022 viral genomes comprised a vast monophyletic group, despite the identification of significant sequence divergence across strains with the emergence of many subclades. This difference is inconsistent with the virus’s rapid diversification over the last several months of the 2022 outbreak. Instead, it represented persistent microevolution since the outbreak in 2017–2018.
Like the most recent common ancestor (MRCA) for the 2017–2018 epidemic, the MRCA for the outbreak in 2022 may be traced back almost 20 years. In addition, the 2022 outbreak MPXVs were more closely linked to strains exported from Africa at the time of the last outbreak than to strains prevalent in Nigeria around that time. Additionally, a strain from an individual traveling to the US from Nigeria in 2021 could also be associated with the source of the outbreak in 2022. Since the outbreak in 2017–2018, the most probable assumption is that MPXV has circulated silently and without detection in many non–MPXV–endemic nations outside of Africa.
Genomic alterations in the MPXV
Multiple genomic alterations were identified in the MPXVs prevalent in 2022. A minimum of 51 single-nucleotide polymorphisms (SNPs) and some larger deletions/insertions differentiated the initial 18 viral genomes observed in the 2022 epidemic from those detected in 2017–2018. Among the 52 SNPs, 26 SNPs resulted in amino acid modifications, whereas 21 were synonymous substitutions. More SNPs were detectable in genomic sequences from 2022, which explained the divergence observed within the epidemic. Future research can help assist in the determination of the phenotypic effects of these alterations, which may be related to mutational pressure and adaptability.
The team also noted that before 2018, viruses from the two MPXV clades had almost comparable amounts of substitution types, and by 2022, the fraction of guanine (G)>adenine (A)/cytosine (C)>thymine (T) transitions within clade II viruses had doubled. Significant substitution alterations presumably reflected the editing ability displayed by the human apolipoprotein-B messenger ribonucleic acid (mRNA) editing enzyme, catalytic subunit 3G (APOBEC3G) enzyme, which catalyzed strand-specific C>uracil (U) deamination and resulted in G>A substitutions in the viral genome complementary strands.
Conclusion
The study findings indicated that the MPXV virus has been silently circulating for almost 20 years, likely in several non-MPXV-endemic nations outside Africa. In addition, the nucleotide substitution pattern demonstrated a distinct genetic hallmark of a recent host alteration. These findings can have significant implications for public health as the shifting epidemiology associated with MPXV infections and the viral circulation among humans in non-MPXV-endemic countries necessitate heightened surveillance.

_6e98296023b34dfabc133638c1ef5d32-620x480.jpg)





;Resize=(1200,627)&impolicy=perceptual&quality=medium&hash=5667f163657ed9a86cb2fe94dc7774a0e26bac9c11c109dea2ebc9588019c1eb)
