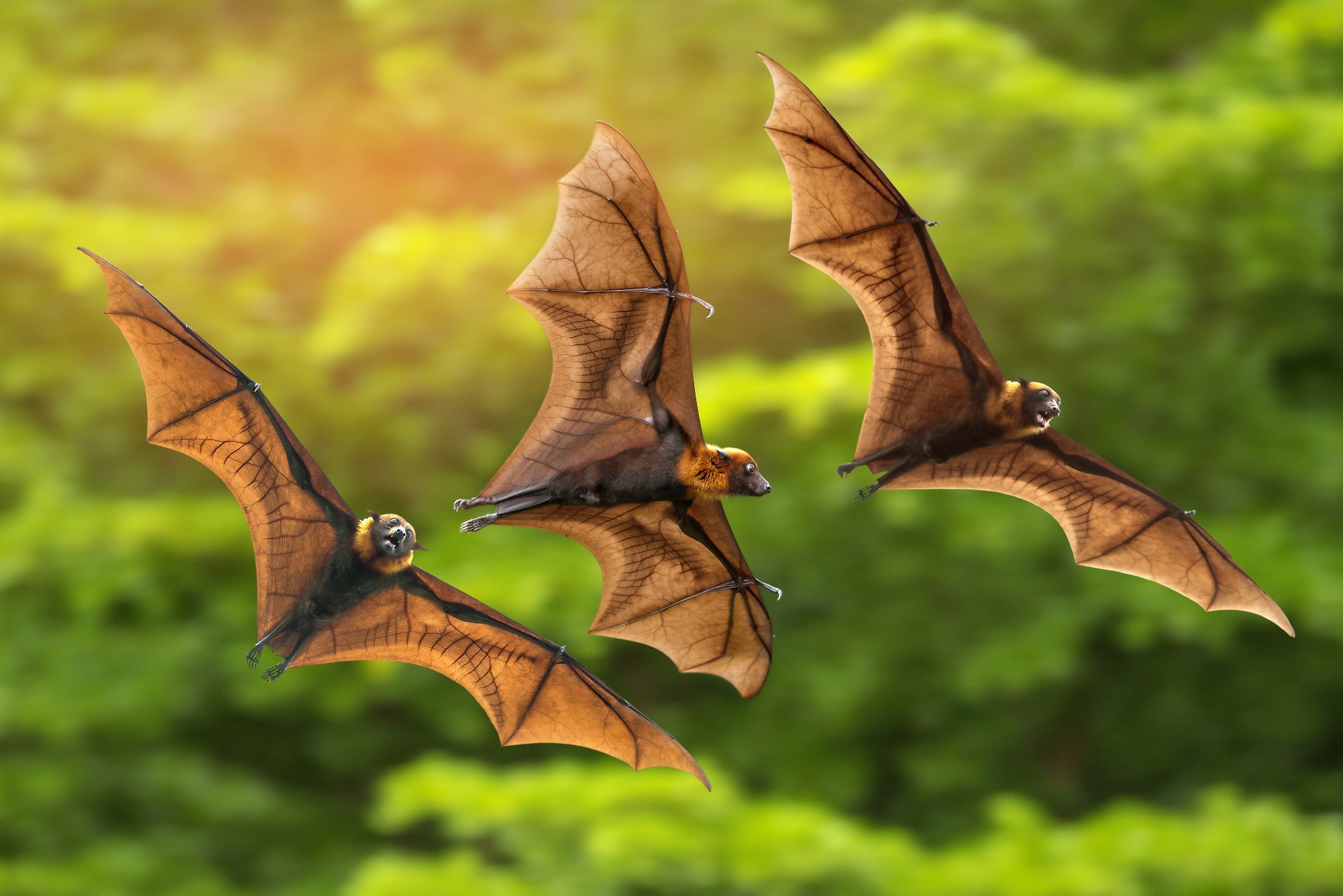
In a recent study published in Emerging Infectious Diseases, researchers performed bat sarbecovirus isolation experiments using Rhinolophus cornutus bats from several regions in Japan that were phylogenetically situated in the same cluster of viruses related to severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2).

Highly pathogenic human beta-coronaviruses (β-CoVs) such as severe acute respiratory syndrome coronavirus (SARS-CoV), SARS-CoV-2, and Middle East respiratory syndrome CoV (MERS-CoV) have been considered to originate from bats. Therefore, bat β-CoV surveillance is essential to improve understanding and to assess the potential of β-CoV spillovers in humans. Sarbecoviruses identified in Asian nations such as China have been reported to be genetically diverse; however, the genetic variations and distribution of bat sarbecoviruses in Japan have not been well-characterized and require further investigation.
The authors of the present study previously showed that a VSV (vesicular stomatitis virus)-based viral pseudotype comprising the S (spike) protein of Rc-o319 sarbecovirus from R. cornutus infected only Rc-ACE2 (angiotensin-converting enzyme 2)-expressing cells, but not human ACE2 (hACE2)-expressing or other Rhinolophus ACE2–expressing cells.
About the study
In the present study, researchers reported on the identification, isolation, and biological and genetic characterization of sarbecoviruses originating from bats in different Japanese locations.
Fecal samples were obtained from the R. ferrumequinum and R. cornutus bat species in Chiba, Shizuoka, and Niigata prefectures. Real-time RT-PCR (reverse transcription polymerase chain reaction) was performed, and the team established RcACE2-expressing Vero cells (Vero-RcACE2) based on Vero/transmembrane protease serine 2 (TMPRSS2) cells.
Next-generation sequencing (NGS) analysis was performed to determine the whole-genome sequence of all viral isolates, and the sequences were deposited in GenBank. In addition, a similarity plot analysis was performed for the full genome sequence by utilizing every isolate as a query. A phylogenetic tree was constructed of bat sarbecoviruses in Japan based on whole-genome sequences with the maximal-likelihood analysis and bootstrapped replicates.
The S RBM (receptor-binding motif) of bat isolates from Japan were aligned with those of other sarbecoviruses. For assessing the growth kinetics of sarbecovirus isolates from bats in Japan, R. cornutus bat isolates Rc-os20, Rc-o319, Rc-mk2, and Rc-kw8 or SARS-CoV-2 (B.1.1.7 strain) were inoculated into Vero-RcACE2 [RcACE2, Vero/TMPRSS2 (WT)], Vero-hACE2 (hACE2), or Vero-ACE2KO (ACE2KO) cells and the viral titers were determined using plaque assays.
Results
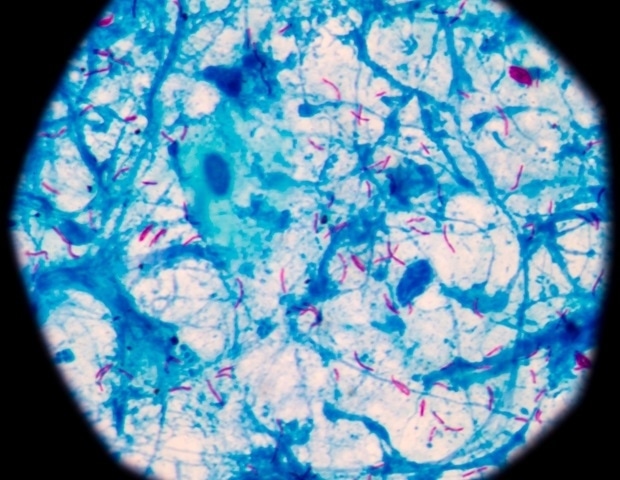
The isolates showed efficient growth in cells that expressed Rhinolophus cornutus ACE2 but did not grow well in hACE2-expressing cells, indicating that the bat isolates had a narrow range of hosts. RT-PCR analysis enabled the successful detection of the envelope (E) gene sequence of sarbecoviruses in one or two Rhinolophus cornutus samples in every prefecture. Bat sarbecoviruses were successfully isolated by utilizing Vero-RcACE2 cell cultures, with enormous cytopathic effects and syncytium formation from RT-PCR‒positive samples of bat feces from every prefecture.
Bat isolates from Shizuoka, Niigata, and Chiba prefectures were designed as Rc-kw8, Rc-os20, and Rc-mk2, respectively. Further, Rc-o319 was isolated by utilizing Vero-RcACE2 cell cultures. Contrastingly, all samples of Rhinolophus ferremuquinum species were negative, indicating that bat sarbecoviruses were distributed among Rhinolophus cornutus bats in Japan. The sequences from all bat strains isolated from Japan were highly homologous (ranging between 95% and 97%). However, Rc-os20 and Rc-mk2 lacked the region coding for ORF8 (open reading frame 8).
The team observed high similarities among the bat isolates throughout the genome sequence, with the exception of sites coding for the S gene receptor-binding domain (RBD) and N-terminal domain (NTD). However, Rc-kw8 and Rc-o319 NTDs were found to be conserved. No significant recombination was observed among the bat isolates. In the phylogenetic analysis, the isolates were observed in one genetic cluster situated in a genetic clade comprising sarbecoviruses related to SARS-CoV-2.
Phylogenetic S RBD analysis findings indicated that the isolates from Japan positioned in the genetic clade of viral organisms utilizing ACE2 ortholog molecules as receptors. Therefore, the team assumed that the novel strains isolated from Japan utilize RcACE2 as receptors. The bat isolates replicated efficiently only in Vero-RcACE2 but not in Vero-ACE2KO, Vero/TMPRSS2, or Vero-hACE2 cells. The findings indicated that the bat isolates depended on RcACE2 for their infectivity.
Contrastingly, SARS-CoV-2 showed efficient replication in Vero-RcACE2, Vero-hACE2 cells, and Vero/TMPRSS2 cells but did not replicate well in Vero-ACE2KO cell cultures, indicating that their infectivity depended on the presence of multiple ACE2 receptors, including those of Rhinolophus cornutus bats. The findings indicated that the bat isolates of Japan only utilized bRcACE2 as receptors.
The majority of genome sequences among strains from Japan were highly conserved, which might be since Rhinolophus spp. bat species migrate to relatively short distances and do not frequently have cross-contact with other groups of bats. The exceptions were S regions coding for RBD and NTD, which showed extensive genetic variation due to immunological pressures, indicating that the bats diverged from the ancestral strain in recent times.
Overall, the study findings showed that the bat isolates utilized batACE2 in the form of receptors with no replication in cells expressing hACE2, therefore, being a characteristic type and indicating scarce human infecting potential. Sarbecoviruses may mutate and lead to human infections through intermediate host species in livestock or wildlife. Therefore, studies of sarbecovirus epidemiology in wild animals, including bats, must be conducted with long-term risk assessments of their potential for zoonotic transmission.